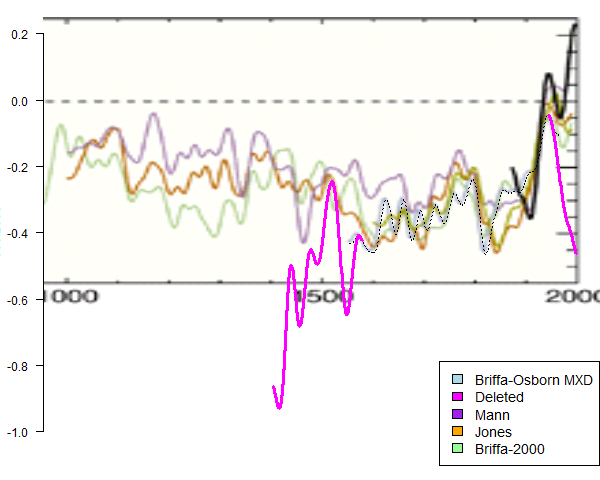
In a comment, Eric Steig suggested direct estimation of temperature trends in West Antarctica(WA) (as opposed to estination via fitted EOFs). For trend estimation, the methods of S09 and O10 have good and bad points. A plus is that they do estimate for a prescribed area. Against that is that it isn't obvious, with variable sea ice, what area to prescribe. A minus is the indirectness. Temperatures are used to fit basis functions (EOFs), and the trend is obtained from the fitted function. Whether through a small number of EOFs, or via regularisation (kgnd), this interpolated step restricts degrees of freedom.
TempLS is designed for the direct estimation of trends via least squares model fitting, so I thought I would try that direct approach. I deferred the project while I looked for a suitable area weighting scheme, and that is now available.
| | Background to this post can be found in earlier discussions here,
here,
here, here,
here, and
here
The discussion is prompted by two papers:
S09, a 2009 Nature paper by Steig et al and RO10 (or O10), a 2010 J Climate paper by O'Donnell et al.
There has been heated blog controversy about these papers - this post is just one entry point. Here is a Real Climate post following S09's original appearance, and here is Eric Steig commenting on O10 (this led to other heated postings).
There are earlier discussions of area weighting by RO10 authors at tAV, eg here and here.
|
The results below broadly confirm the earlier spatial analyses, and follow the regional breakdown of S09 and O10, though often with higher trends. The continent ground station trend, 1958 to 2009 and not area weighted, was 0.175 ± 0.03 °C/decade. This is consistent with my earlier EOF-based result, and higher than S09 or O10. Area weighting brought it down to 0.114 ± 0.035 °C/decade, similar to S09 (with AVNRR). Adding equally weighted AVHRR pushed it up to 0.137 ± 0.037 °C/decade. I think these results add a degree of confirmation to those earlier analyses, and to my EOF-based analysis, and tend to confirm a positive warming trend, especially outside East Antarctica.
Something that needed attention is the regional tagging of the data I am using, which comes from the data Ryan put
on line, and earlier from S09. There are four regions, Peninsula, WA, Ross Sea, EA. But the satellite tags are bounded inconsistently with the ground stations, so I retagged them as shown in this map. Ground stations are shown with large dots, and the subset of AVHRR stations (described
here) are small dots, both colored by region.

That done, I have computed trends for these regions and the continent using the
OLS methods of TempLS V1. As with some earlier posts, I have mixed AVHRR and ground stations, with various relative weightings (including ground only).
Continent ground station trends
These are difficulties with the sparse samples in some tof the regions, most notably with West Antarctica. So the whole continent is simplest. I'll start with unweighted (by area):

Stations reporting in each year | 
A map of ground stations colored by region
|
 | |
The trend is quite high at 0.175°C/decade. Now I'll add weighting by area, as described
here. The map shows for a specific month, Dec 1987, the triangular mesh created, the subdivision used to weight each station, and the circles show, by area, the size of the weights. I have not smoothed, nor tried to cut out the Weddell Sea.
The trend is lower, at 0.114°C/decade. This is in line with earlier observations that area weighting reduces the trend, probably by boosting the effect of the less rapidly warming East Antarctica stations, which occupy the largest region.
Adding AVHRR readings
As with
previous posts, I've introduced the satellite AVHRR data, which comes as grid data, as artificial stations. I've selected only 1 in 4, so there are 1392 of them. A factor is needed to determine the relative weighting for combining them with ground stations. I now use a range from 0 (all satellite) to 1 (all ground), with 0.5 indicating that total weights for satellite and ground are equal. That is for all years, so that before 1980, there is ground data only, and the factor doesn't matter. Because of the way it is totalled, this means sat data gets a higher weighting (at 0.5) after 1980. This is a small effect.
So here are results from satellite data only. Like O10, I'm using Eric Steig's archived AVHRR data, which goes up to 2006 only.
As expected, the trend is quite high. One of the reasons for interest in S09 was that it looked into the discrepancy between the ground trend (lowish) and AVHRR (high).
So now looking at the combination of satellite and ground, remembering that before 1980 it is ground only:
So the trend is lower - closer to ground than AVHRR, reflecting the shorter period of AVHRR.
Antarctic Peninsula trends
So now I look at regional trends. I'll start with the Peninsula, which does not have problems of years without data. Here are ground stations only:
AS expected, the trend is high. Now if the AVHRR stations are included, with equal (total) weighting:
East Antarctica trends
Again firstly the ground stations:
As reflected in work in previous posts (and S09, O10) ground stations in East Antarctica have been warming very slowly. The inclusion of AVHRR results at nominally equal weighting increases this by a fairly small amount:
West Antarctica and Ross Sea
I found a specific difficulty with WA, which I think Ryan and Eric have
mentioned. In about 1980, about the time AVHRR satellite information is
available, there was also a change to measurement (AWS). Only one
station in WA was reporting before then, Byrd, and there was a gap of
four years with no readings at all. Byrd changed to AWS, and this is
listed as a new station. So there is no way of bridging the gap without
making some assumption about how the old Byrd relates to Byrd AWS. And even that would be slender information.
A first option for WA is to combine it with the Ross Sea stations, where there is a fuzzy boundary anyway. There are just enough stations pre-1980 to get a result:
Again, as expected, the trend is high. Adding AVHRR stations does not change this.
West Antarctica trends
The other option for WA is to look at results after 1980 only. That gives:
The trend is very high indeed, but so is the uncertainty, and the stations are few.
It seems the AVHRR data stabilizes the average when it is available, so just checking those years:
This did reduce the trend slightly, and improve the error range.